Since DNA’s structure was discovered in 1953, numerous techniques have been developed to use this knowledge to learn more about how living things function and solve genetic questions. DNA (Deoxyribonucleic Acid) is the genetic material that contains the “code of life.” Each person has a unique DNA code in each of their cells that can be used to identify them and their children. We get our unique DNA code from our biological parents in equal amounts.
DNA Science Explained
Paternity testing and other similar DNA tests are the gold standard for identifying biological relationships.
Everything You Need to Know about how DNA Science Works

DNA Biology: Structure and Replication
The building blocks of DNA are known as nucleotides (abbreviated A, T, C, and G), and appear in DNA as letter pairs. There are about 3 billion letter pairs in our DNA code.
In humans, most of the DNA is in the form of tightly-coiled strands called chromosomes, found inside the cell nucleus. There are 46 chromosomes in a human cell arranged in 22 autosomal (non-sex chromosome) pairs and two sex chromosomes (XY for males and XX for females). If you unwind each chromosome and place them end-to-end, you’ll have a long, double-stranded DNA helix that is about 3 meters in length—all from one microscopic human cell.
The DNA helix looks like a twisted ladder or spiral staircase. The “steps” are composed of the four bases: adenine (A), thymine (T), guanine (G), and cytosine (C), and are held together by hydrogen bonds that connect specific pairs of these molecules together: A–T and G–C. The arrangement of these molecules, called the DNA sequence, spells out the instructions for our physical characteristics and body functions. These instructions are found in units called genes.
The specific pairing of DNA molecules presents a simple mechanism for their replication. Replication occurs whenever a cell divides in two during growth and development. During replication, the helix unwinds and an enzyme separates the two strands. Another enzyme, DNA polymerase, adds the four molecules A, T, C, and G to each strand according to the strand’s sequence: A is added opposite a T on the strand, and C is added opposite a G. This “complementary base pairing” allows the replication to occur quickly and reliably.
The products of replication are two sets of double-stranded DNA molecules that have exactly the same sequence as the original. During cell division, each cell receives one set of DNA. In this way, all the cells of the body have the same DNA molecules.
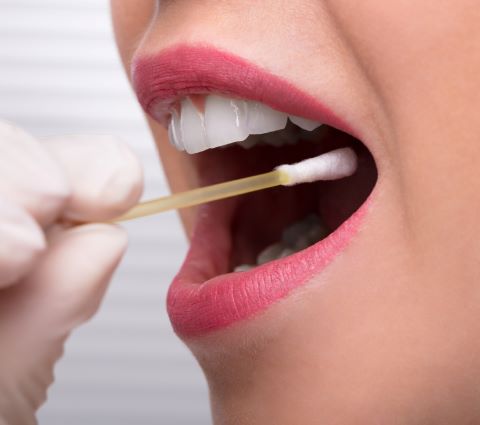
DNA Extraction
DNA is found in the nucleus of a cell, which is a lot like the yolk of an egg. DNA is extracted from a cheek (buccal) swab sample by breaking open the cells and separating the DNA from the other parts of the cell. DDC uses scientifically-accepted methods to perform the DNA extraction step, along with robotic instruments for precision and accuracy.

The Polymerase Chain Reaction
Polymerase Chain Reaction or PCR, is a technique that allows the generation of millions of copies of targeted areas (loci) in the DNA. PCR harnesses the cell’s ability to replicate DNA before cell division, but in a test tube. PCR is used commonly in the scientific community to perform a number of different tests and is used by DDC for paternity testing.
Once the DNA is extracted, it is combined with chemicals to support the PCR reaction, including primers. Primers are small pieces of synthetic DNA labeled with florescent tags that flank the region to be analyzed. The DNA building blocks (A, T, C, G) and an enzyme to support the replication of the DNA (DNA polymerase) are included in the mixture.
For paternity testing, DDC uses a multiplex PCR reaction that tests 21 different locations (loci) in the DNA at once. This is accomplished using an instrument known as a thermocycler that heats and cools the DNA. Each round of heating and cooling doubles the amount of DNA at the target locations, so that after 28 cycles of heating and cooling over a million copies of each starting DNA are present.

Short Tandem Repeat (STR) Analysis
STR loci are used for relationship analysis, including paternity, maternity, and siblingship testing. STR loci contain a 4-letter DNA repeat and the number of repeats can vary. There are two copies of each of the STR loci. For relationship testing DDC examines 20 different STR loci plus a locus that indicates gender.
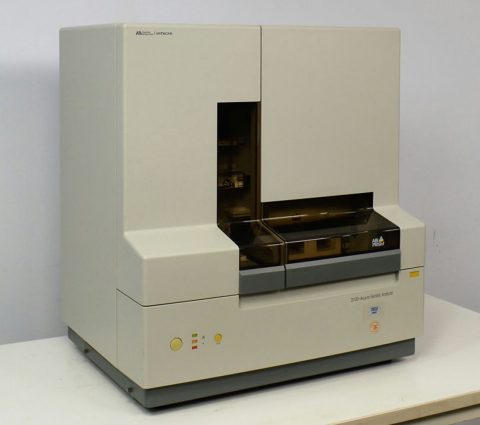

Genetic Analyzer
The number of repeats at each STR location is determined by analyzing the products of the PCR reactions with a Genetic Analyzer. These instruments allow the analysis of up to 48 samples simultaneously. The products of the PCR are loaded into a small capillary tube and an electrical current is applied, which moves the DNA down the tube in a size- dependent fashion. This is so that the smaller DNA fragments “win the race” to the end of the tube. As the DNA fragments exit the tube, a laser interrogates them and the florescent tags incorporated during PCR are activated. The instrument captures this information and determines the size of the fragment by the time it took to exit the tube.

Paternity-Test Report
The information from the Genetic Analyzer is incorporated into the final report. At each STR locus, the number of repeats detected is listed as a number, i.e., a 12 means there are 12 four-letter repeats present. Each of the numbers represents an allele, or an alternate form of the DNA at that location. Since each STR locus is assigned to a chromosome position and chromosomes are present in pairs, there are typically two numbers per person at each STR location (heterozygous).
When only one number is shown at a locus, both chromosomes in the pair have the same number of repeats (homozygous). A parent and child must share a common allele (number) to be the biological parent. Once the results of all 20 STR locations are evaluated, a calculation is performed to provide the numerical support for the tested biological relationship and is expressed as the probability of relationship. Learn more about how to understand a paternity report here.

Solving Crimes and Mysteries
A large portion of human DNA does not code for genes (units of DNA that contain instructions for creating proteins). These non-coding regions nevertheless contain much information useful for forensic and other human identification applications. Non-coding regions of DNA contain many of the markers used for identity testing.
- Short Tandem Repeats (STRs) as discussed above are useful for human identification. The combination of STR alleles is unique to each person and can be used to either include or exclude a person as the source of a biological forensic evidentiary sample. These DNA profiles or DNA fingerprints, are routinely used to solve crimes.
- Single Nucleotide Polymorphisms (SNPs) are individual letter changes in the DNA, i.e. an A in one person and a G at the same position in another person. There are millions of such changes in our DNA and they can be used for identification and ancestry testing. DDC’s GPS Origins™ test uses about 800,000 SNPs to determine ancestry and migration routes of our ancestors.
- Mitochondrial DNA (mtDNA) is found in the energy-producing organelles of the cell called the mitochondria. Most of our DNA is found in the nucleus of cells in two copies; one from each biological parent. Mitochondrial DNA, on the other hand, is a shorter piece of DNA found in hundreds or even thousands of copies per cell and is inherited from the biological mother only.
Mitochondrial DNA analysis is performed by determining the DNA sequence of a specific region of the mitochondrial DNA known as the hypervariable region. Although not unique individually, mitochondrial DNA can help establish a maternal lineage, since the mitochondrial DNA is inherited only from the biological mother.
Non-coding regions in mtDNA, called hypervariable regions, have significant sequence differences that can be used to differentiate two unrelated individuals. In contrast, biological relatives with a common maternal ancestry have the exact or very similar sequences at this region. Mitochondrial DNA is inherited through the mother, and the biological relationship of two or more individuals may be determined by examining their mtDNA sequences at the hypervariable regions.
Mitochondrial DNA testing was used to identify the bodies of the Romanovs found in a burial site in Yekaterinburg, Siberia. Mitochondrial DNA sequences from a maternal relative, Prince Philip, matched the DNA extracted from the remains of the Tsarina and her daughters; mtDNA sequence from the Duke of Fife (a maternal relative of the Tsar) was used to confirm the identity of the Tsar’s remains. - Y chromosome typing (Y-STR) uses short tandem repeats found on the Y chromosome to trace the ancestry of males through the paternal line. The Y chromosome is only passed from father to son, relatively unchanged through the generations, and thus males related through a paternal line share the same Y-STR pattern. While Y-STR analysis gives clues about recent male ancestry, another type of markers called SNPs (pronounced “snips,” single nucleotide polymorphisms) give clues about deep male ancestry—the early origins of the different human populations, going back even up to 100,000 years. Analysis of SNPs combined with Y-STR typing is used to recreate historical migrations of waves of populations across the world.
DNA Testing With Reputable Accreditations












Accreditations apply to legal DNA tests under scope.
Why Choose DDC for Paternity Testing?
- Trusted Expertise: For decades, customers in Boston and beyond have trusted us with over 20 million DNA tests.
- Convenient Location: Visit one of our nearby locations in Boston or schedule DNA collection at a location near you.
- Free Consultations: Not sure which test is right for you? Our DNA experts are here to help.
- Fast Results: With same-day paternity testing options available, get the answers you need quickly and reliably.
DNA Science is our Life
Our 11 in-house PhD geneticists along with a staff of highly-trained and qualified laboratory technicians and support personnel keep up with the very latest DNA technology and techniques, so you can be sure your test is in good hands. It’s our passion.
Don’t wait to get the answers you need.
DDC can provide clarity and peace of mind for your most important family questions.
Se Habla Español


